What is transcriptomics?
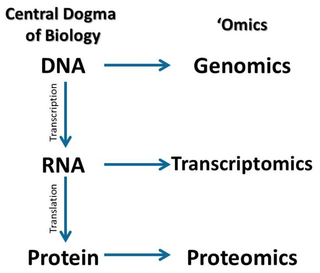
Transcriptomics is the study of the transcriptome, which is the complete set of RNA transcripts that are produced by the genome, under specific circumstances or in a specific cell. [1] Transcriptomics uses two main methods which are RNA sequencing and microarray analysis. Comparing various transcriptomes allows the identification of genes that differentiate in expression in distinct cell populations, or in response to different treatments. [1]
What is RNA Sequencing?
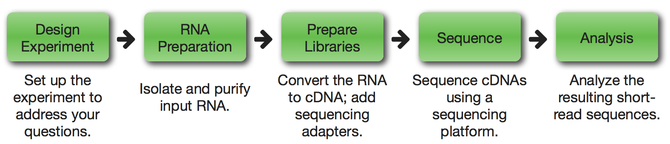
An RNA sequencing (RNA-seq) experiment involves making a collection of cDNA fragments matched by specific constant sequences (adaptors) that are necessary for sequencing. This collection, often called a library, is then sequenced using short-read sequencing. This sequencing produces millions of short sequence reads that correspond to individual cDNA fragments. [2]
What is Microarray Analysis?
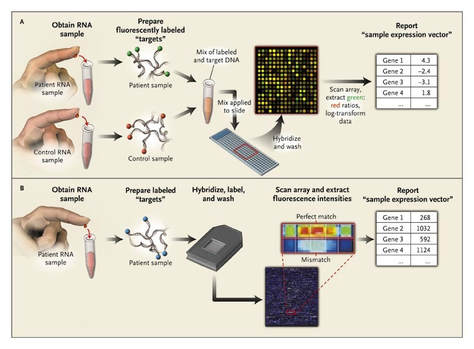 Figure 3. Experimental Layout of Microarray Analysis
Figure 3. Experimental Layout of Microarray Analysis
In order to perform microarray analysis, mRNA molecules are collected from experimental and reference samples. The samples are converted into complementary DNA (cDNA) and each sample is labeled with a fluorescent probe with different colors. The samples are then mixed and allowed to bind to the microarray slide. The microarray is then scanned to measure the expression of each gene printed on the slide. [3] The spots expressed on the microarray scan represent which genes are expressed and their levels of expression.
How can transcriptomics be used to study TCF4?
Comparing transcriptomes in different cell types can provide important information regarding the mechanisms behind biological pathways or processes. For example, in this study, I am interested in observing the effects of TCF4 activation in cell differentiation of lung and heart tissue cells. Knock-out of TCF4 in mice and then comparing the transcriptomes of these mutants to the wild type mice transcriptomes can provide important information, This information can identify important proteins or functions that contribute to the processes of cell differentiation in lung and heart tissue.
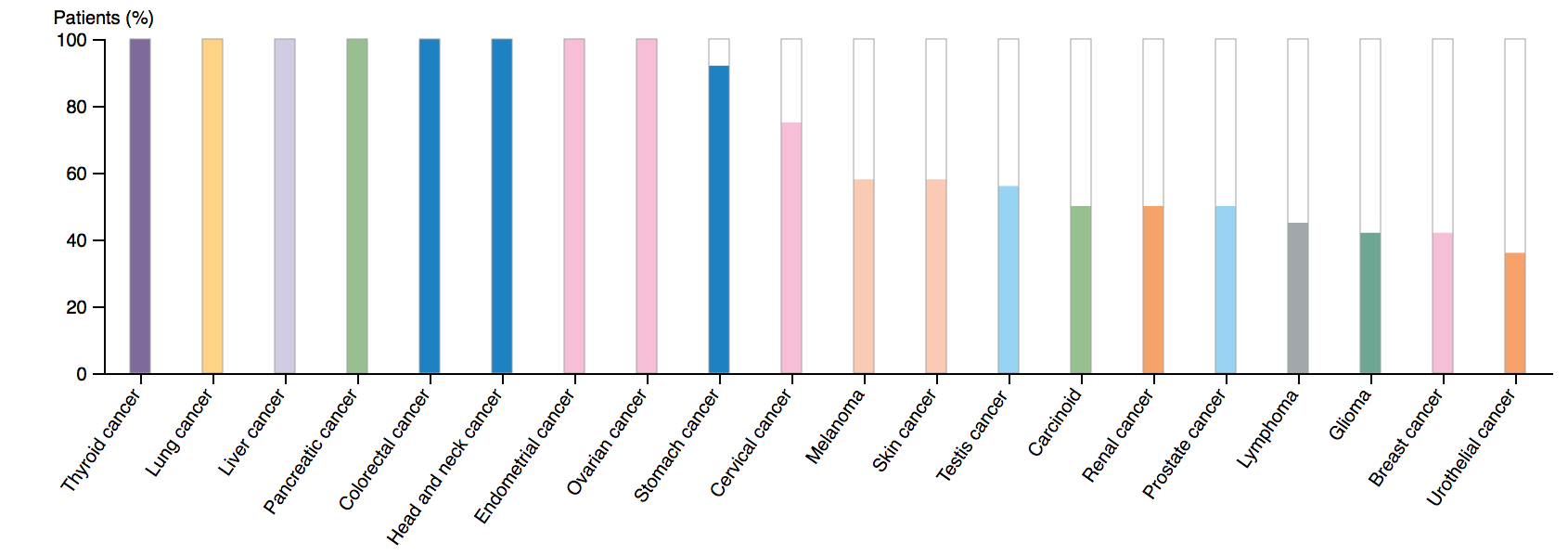
The Human Protein Atlas (HPA) is a program that maps human proteins in cells, tissues and organs using various methods and techniques including transcriptomics. HPA provides information regarding cell localization, gene descriptions, protein expression, cell function, gene ontology, and levels of gene expression in various tissues. Using HPA, I was able to identify the role of TCF4 gene expression in various tissue type cells as well as its role in cancer development in different tissue types. One interesting finding was that although TCF4 expression is highest in the brain, its contribution to cancers of the brain (glioma) is extremely average. However, TCF4 gene expression is high in ovarian, stomach, liver, lung, endometrial and thyroid cancers. This seems contradictory, considering that TCF4 plays such an important part in neurological development and neuronal cell differentiation. This information suggests that TCF4 plays an important role in cell differentiation processes in other tissue types such as lung tissue. It also emphasizes how TCF4 is very broadly expressed across various different tissue types.
The Human Protein Atlas (HPA) is a program that maps human proteins in cells, tissues and organs using various methods and techniques including transcriptomics. HPA provides information regarding cell localization, gene descriptions, protein expression, cell function, gene ontology, and levels of gene expression in various tissues. Using HPA, I was able to identify the role of TCF4 gene expression in various tissue type cells as well as its role in cancer development in different tissue types. One interesting finding was that although TCF4 expression is highest in the brain, its contribution to cancers of the brain (glioma) is extremely average. However, TCF4 gene expression is high in ovarian, stomach, liver, lung, endometrial and thyroid cancers. This seems contradictory, considering that TCF4 plays such an important part in neurological development and neuronal cell differentiation. This information suggests that TCF4 plays an important role in cell differentiation processes in other tissue types such as lung tissue. It also emphasizes how TCF4 is very broadly expressed across various different tissue types.
Conclusion
Transcriptomics is a crucial aspect of genomics due to its ability to transform transcriptomes into informative tools of genetic research. Comparing transcriptomes across various tissue types or proteins can reveal ground-breaking information regarding the complex pathways of a particular process. For example, when studying TCF4, it would be helpful to compare transcriptomes of wild type mice to TCF4 knockout mutant mice in order to observe the effects found in heart and lung tissue, specifically their cell differentiation processes. This information can be used to identify important proteins or key molecular players that are involved in TCF4's role as a transcription factor and E-protein, in regards to cell differentiation.
References
Header: http://www.mfpl.ac.at/rna-biology/wp-content/uploads/2012/03/RNA-BIOLOGY-WALLPAPER.png
[1] Nature. (n.d.). Transcriptomics. Retrieved March 8, 2018, from https://www.nature.com/subjects/transcriptomics
[2] University of Oregon. (n.d.). RNA-seqlopedia. Retrieved from https://rnaseq.uoregon.edu
[3] Nature.com. (n.d.). Microarray. Retrieved March 8, 2018, from https://www.nature.com/scitable/definition/microarray-202
Figure 1. https://wildlifesnpits.wordpress.com/2015/11/18/transcriptomics-for-conservation/G
Figure 2. https://rnaseq.uoregon.edu
Figure 3. http://www.nejm.org/doi/full/10.1056/NEJMra042342
Figure 4. http://www.proteinatlas.org/ENSG00000196628-TCF4/pathology
Header: http://www.mfpl.ac.at/rna-biology/wp-content/uploads/2012/03/RNA-BIOLOGY-WALLPAPER.png
[1] Nature. (n.d.). Transcriptomics. Retrieved March 8, 2018, from https://www.nature.com/subjects/transcriptomics
[2] University of Oregon. (n.d.). RNA-seqlopedia. Retrieved from https://rnaseq.uoregon.edu
[3] Nature.com. (n.d.). Microarray. Retrieved March 8, 2018, from https://www.nature.com/scitable/definition/microarray-202
Figure 1. https://wildlifesnpits.wordpress.com/2015/11/18/transcriptomics-for-conservation/G
Figure 2. https://rnaseq.uoregon.edu
Figure 3. http://www.nejm.org/doi/full/10.1056/NEJMra042342
Figure 4. http://www.proteinatlas.org/ENSG00000196628-TCF4/pathology
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.