Pitt Hopkins Syndrome & TCF4 in Intellectual Disability
Pitt Hopkins Syndrome (PTHS) is a rare genetic disease with only 50 cases reported worldwide. [1] It is associated with symptoms including epilepsy, lack of speech, low muscle tone and breathing problems. [2] The cause of PTHS is haploinsufficiency of the TCF4 gene. This indicates that individuals with PTHS have only one functional copy of TCF4 and that this copy is not sufficient to produce enough gene product. In nerve cells, this haploinsufficiency results in the inhibition of cell differentiation which contributes to the occurrence of symptoms that impact the nervous system's development and function.
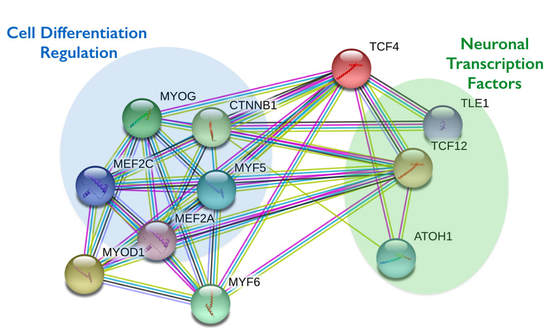
TCF4 is most closely related to the processes of cell differentiation in addition to apoptosis. It is active in the cytosol and nucleoplasm of cells and acts as both a transcription factor and E-protein. [3] Since TCF4 is dependent on its interactions with other proteins to serve its function, its protein-protein interactions are definitive of its own function. TCF4 has one protein domain, which is a basic helix-loop-helix domain, which is a large family of transcription factors. This protein domain is crucial for development of the nervous system and muscle. [4] The protein interaction network for TCF4 reveals high association with proteins involved in cell differentiation and neuronal transcription factors. [Figure 1] These are expected due to TCF4's association with cell differentiation and the effect of TCF4 haploinsufficiency on the nervous system.
Overall, TCF4 is well conserved among a wide variety of species. However, for the purposes of this study, Mus musculus (the house mouse) was most appropriate. There were various reasons as to why mice were the most appropriate model organisms for this study. Two of the more general reasons why mice are used as model organisms in research is due to their cost-efficiency and time-efficiency. Mice are significantly cheaper to maintain than some other model organisms and have shorter generation times, making them easier to use for research. In addition, their similarities to humans in terms of muscular structure and function make them a great model organism. Mice are often used to study human muscle diseases and muscle development. While reviewing past literature, mice were also used in the majority of studies pertaining to TCF4. Their tissue can also be analyzed for the PTHS phenotype of low muscle tone via cyrosectioning. [Figure 1] Cyrosectioning is the use of liquid nitrogen to freeze tissue samples to use for rapid microscopic analysis.
TCF4 is most closely related to the processes of cell differentiation in addition to apoptosis. It is active in the cytosol and nucleoplasm of cells and acts as both a transcription factor and E-protein. [3] Since TCF4 is dependent on its interactions with other proteins to serve its function, its protein-protein interactions are definitive of its own function. TCF4 has one protein domain, which is a basic helix-loop-helix domain, which is a large family of transcription factors. This protein domain is crucial for development of the nervous system and muscle. [4] The protein interaction network for TCF4 reveals high association with proteins involved in cell differentiation and neuronal transcription factors. [Figure 1] These are expected due to TCF4's association with cell differentiation and the effect of TCF4 haploinsufficiency on the nervous system.
Overall, TCF4 is well conserved among a wide variety of species. However, for the purposes of this study, Mus musculus (the house mouse) was most appropriate. There were various reasons as to why mice were the most appropriate model organisms for this study. Two of the more general reasons why mice are used as model organisms in research is due to their cost-efficiency and time-efficiency. Mice are significantly cheaper to maintain than some other model organisms and have shorter generation times, making them easier to use for research. In addition, their similarities to humans in terms of muscular structure and function make them a great model organism. Mice are often used to study human muscle diseases and muscle development. While reviewing past literature, mice were also used in the majority of studies pertaining to TCF4. Their tissue can also be analyzed for the PTHS phenotype of low muscle tone via cyrosectioning. [Figure 1] Cyrosectioning is the use of liquid nitrogen to freeze tissue samples to use for rapid microscopic analysis.
TCF4 haploinsufficiency is known to disrupt cell differentiation in neural crest cells. Yet, the mechanisms of how TCF4 functions in muscle tissue development are unclear. My primary goal is to determine if TCF4 is required for the differentiation of muscle tissue cells during embryonic development. My objective is to explore how TCF4 mediates cell differentiation during muscle tissue development. My hypothesis is that deletion of the TCF4 will inhibit the process of cell differentiation in cells found in muscle tissue.
Aim #1: Identify TCF4 mutations that affect cell differentiation processes in muscle tissue
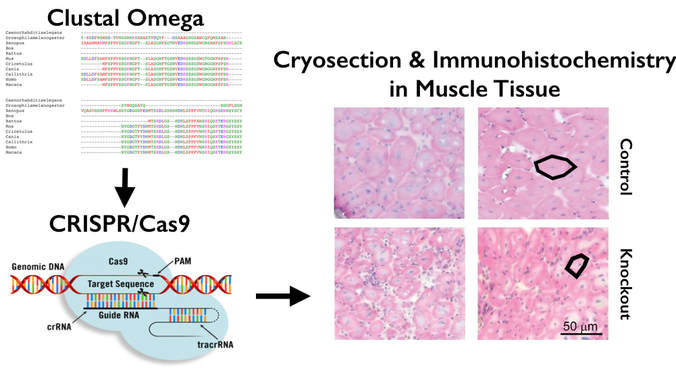
In order to identify TCF4 mutations that affect cell differentiation processes in muscle tissue, I first began with using BLAST to gather FASTA amino acid sequences for homologs of TCF4. Then, I used Clustal Omega for Multiple Sequence Alignment in order to determine the well-conserved amino acid regions for TCF4. After this, I used CRISPR Cas9 to create TCF4 mutant knockout mice. These were then screened for the low muscle tone phenotype found in patients with PTHS. This was done through the use of cyrosectioning and immunohistochemistry. [Figure 2]
The rationale behind this approach is that this workflow will allow me to confirm that cell differentiation processes are related to TCF4 knockout in muscle tissue cells. I hypothesize that the mutant mice will result in lower muscle tone, meaning that the well-conserved regions are associated with muscle cell development and differentiation processes.
The rationale behind this approach is that this workflow will allow me to confirm that cell differentiation processes are related to TCF4 knockout in muscle tissue cells. I hypothesize that the mutant mice will result in lower muscle tone, meaning that the well-conserved regions are associated with muscle cell development and differentiation processes.
Aim #2: Identify new genes important for muscle development in TCF4 mutants
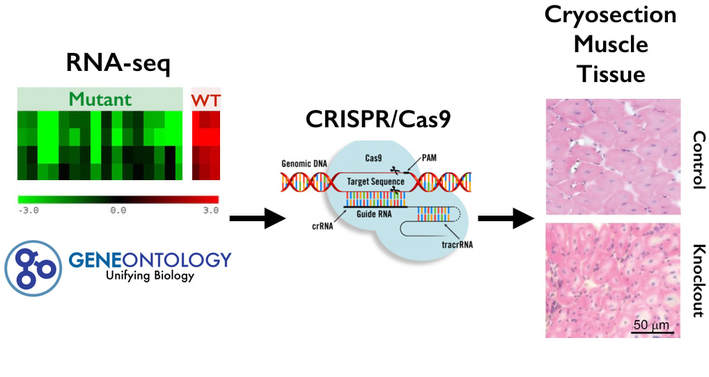
To identify new genes important for muscle development in TCF4 mutants, RNA-seq was used to identify differentially expressed genes using muscle tissue cells from wild type and TCF4 mutant mice. The differentially expressed genes will be sorted using Gene Ontology (GO) to determine which biological processes are most affected by TCF4 knockout. After this, CRISPR Cas9 will be used to create TCF4 mutant knockout mice and to validate these genes by screening once again for the low muscle tone phenotype. [Figure 3[
The rationale behind this approach is that by evaluating expression levels of these genes, I will be able to identify genes affected by the lack of TCF4. Their GO terms will reveal if they are related to muscle tissue cell differentiation processes and muscle function. I hypothesize that TCF4 knockout will result in differentially expression genes crucial for cell differentiation and cell fate determination of muscle tissue. In addition, these genes will have lower levels of RNA expression in the TCF4 mutant mice.
The rationale behind this approach is that by evaluating expression levels of these genes, I will be able to identify genes affected by the lack of TCF4. Their GO terms will reveal if they are related to muscle tissue cell differentiation processes and muscle function. I hypothesize that TCF4 knockout will result in differentially expression genes crucial for cell differentiation and cell fate determination of muscle tissue. In addition, these genes will have lower levels of RNA expression in the TCF4 mutant mice.
Aim #3: Identify new TCF4 interacting proteins involved in muscle tissue development
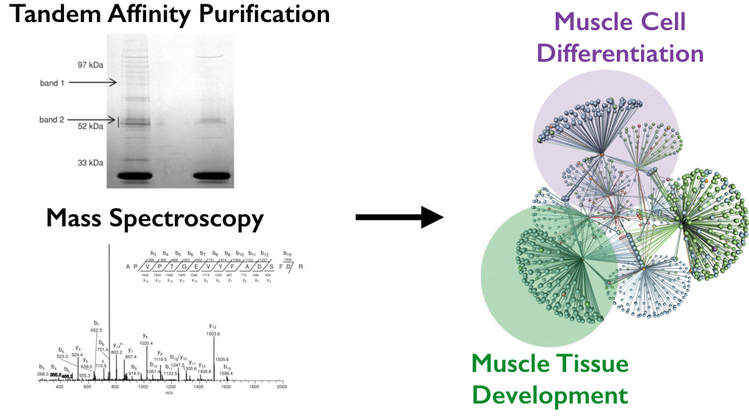
To identify new TCF4 interacting proteins involved in muscle tissue development, I will use co-immunoprecipiation (Tandem Affinity Purification) and Mass Spectroscopy in order to identify new proteins involved in muscle tissue development. They will then be sorted according to GO terms specifically for terms related to muscle cell differentiation and muscle tissue development. [Figure 4]
The rationale behind this approach is that identifying new proteins that interact with TCF4 can reveal new roles specifically in muscle tissue development and cell differentiation. I hypothesize that TCF4 will be identified as a transcription factor that may regulate proteins involved in muscle tissue development.
The rationale behind this approach is that identifying new proteins that interact with TCF4 can reveal new roles specifically in muscle tissue development and cell differentiation. I hypothesize that TCF4 will be identified as a transcription factor that may regulate proteins involved in muscle tissue development.
Future Directions
TCF4 is expressed in a wide variety of tissue types. Hence, conducting similar studies with the use of genomic and proteomic approaches to study TCF4 expression in other tissue types can reveal additional information about its role in biological processes. This could lead to further information regarding possible medicinal approaches or cures for diseases such as Pitt Hopkins syndrome.
Final Talk Draft One
| delacruz4-3-18finaltalkdraft.key | |
| File Size: | 7232 kb |
| File Type: | key |
Final Talk
| delacruz4-25-18finaltalkdraft.pdf | |
| File Size: | 4498 kb |
| File Type: | |
References
Header: https://www.executivechronicles.com/time-to-recognize-intellectual-property-rights/
[1] Peippo, M., & Ignatius, J. (2012, April). Pitt-Hopkins Syndrome.<https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3366706/>
[2] What is Pitt Hopkins syndrome? (2017, October 18). <https://pitthopkins.org/what-is-pitt-hopkins-syndrome/>
[3] Pitt-Hopkins syndrome - Genetics Home Reference. (n.d.). <https://ghr.nlm.nih.gov/condition/pitt-hopkins-syndrome>
[4] Rosenfeld JA, Leppig K, Ballif BC, Thiese H, Erdie-Lalena C, et al. Genotype-phenotype analysis of TCF4 mutations causing Pitt-Hopkins syndrome shows increased seizure activity with missense mutations. Genet Med. 2009;11:797–805.
Figures
http://www.executivechronicles.com/time-to-recognize-intellectual-property-rights/
http://pluspng.com/png-82614.html
https://www.webmd.com/women/ss/slideshow-thyroid-symptoms-and-solutions
https://www.homenaturalcures.com/bone-marrow-home-remedy/
https://www.webmd.com/men/lymph-nodes-in-the-male-retroperitoneum-and-pelvis
https://medlineplus.gov/ency/imagepages/19841.htm
https://www.pancan.org/facing-pancreatic-cancer/about-pancreatic-cancer/what-is-the-pancreas/
https://en.wikipedia.org/wiki/Gastrointestinal_tract#/media/File:Blausen_0432_GastroIntestinalSystem.png
http://www.thehealthsite.com/diseases-conditions/revealed-14-hidden-symptoms-of-kidney-disease/
http://www.wisegeek.org/what-is-an-enlarged-bladder.htm#
https://www.bioscience.co.uk/products/crisprcas9-genome-engineering
https://www.researchgate.net/figure/Tri-lineage-differentiation-potential-of-tissue-cells-Cells-from-bone-marrow-BM-A-C_fig4_259200292
https://www.researchgate.net/figure/Results-of-three-step-validation-of-RNAseq-differential-gene-expression-analysis-Each_fig5_282072768
https://s3.amazonaws.com/go-public/image/go-logo.large.png
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4752115/
https://upload.wikimedia.org/wikipedia/commons/5/53/Multiplex_Human_HIV-1_protein-protein_interaction_network_%28edge-colored_visualization%29.png
Header: https://www.executivechronicles.com/time-to-recognize-intellectual-property-rights/
[1] Peippo, M., & Ignatius, J. (2012, April). Pitt-Hopkins Syndrome.<https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3366706/>
[2] What is Pitt Hopkins syndrome? (2017, October 18). <https://pitthopkins.org/what-is-pitt-hopkins-syndrome/>
[3] Pitt-Hopkins syndrome - Genetics Home Reference. (n.d.). <https://ghr.nlm.nih.gov/condition/pitt-hopkins-syndrome>
[4] Rosenfeld JA, Leppig K, Ballif BC, Thiese H, Erdie-Lalena C, et al. Genotype-phenotype analysis of TCF4 mutations causing Pitt-Hopkins syndrome shows increased seizure activity with missense mutations. Genet Med. 2009;11:797–805.
Figures
http://www.executivechronicles.com/time-to-recognize-intellectual-property-rights/
http://pluspng.com/png-82614.html
https://www.webmd.com/women/ss/slideshow-thyroid-symptoms-and-solutions
https://www.homenaturalcures.com/bone-marrow-home-remedy/
https://www.webmd.com/men/lymph-nodes-in-the-male-retroperitoneum-and-pelvis
https://medlineplus.gov/ency/imagepages/19841.htm
https://www.pancan.org/facing-pancreatic-cancer/about-pancreatic-cancer/what-is-the-pancreas/
https://en.wikipedia.org/wiki/Gastrointestinal_tract#/media/File:Blausen_0432_GastroIntestinalSystem.png
http://www.thehealthsite.com/diseases-conditions/revealed-14-hidden-symptoms-of-kidney-disease/
http://www.wisegeek.org/what-is-an-enlarged-bladder.htm#
https://www.bioscience.co.uk/products/crisprcas9-genome-engineering
https://www.researchgate.net/figure/Tri-lineage-differentiation-potential-of-tissue-cells-Cells-from-bone-marrow-BM-A-C_fig4_259200292
https://www.researchgate.net/figure/Results-of-three-step-validation-of-RNAseq-differential-gene-expression-analysis-Each_fig5_282072768
https://s3.amazonaws.com/go-public/image/go-logo.large.png
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4752115/
https://upload.wikimedia.org/wikipedia/commons/5/53/Multiplex_Human_HIV-1_protein-protein_interaction_network_%28edge-colored_visualization%29.png
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.