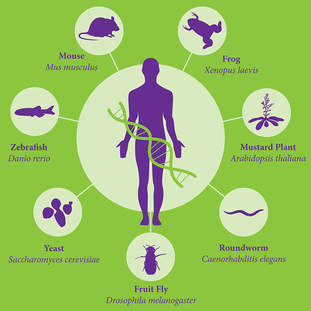
What are model organisms?
Model organisms are species which have been widely studied because they are easy to maintain and breed in a laboratory setting and has particular experimental advantages. [1] They are typically used to help scientists understand biological processes. They are extremely helpful in genetics research for various reasons such as the fact that they breed in large numbers with very short generation times. [1] There are many factors considered when choosing appropriate model organisms such as the biological processes involving the particular gene of interest.
How are model organisms chosen?
There are various scientific and practical factors considered when selecting appropriate species, including [2]:
- Which species will yield the most scientifically accurate and interpretable results?
- According to critical review of the scientific literature, which species have provided the best, most applicable historical data?
- On which species will data from the proposed experiments be most relevant and useful to present and future investigators?
- Which species have special biologic or behavioral characteristics that make them most suitable for the planned studies?
- Which species have features that render them inappropriate for the planned studies?
- Which species present the fewest or least severe biologic hazards to the research team?
- Which species require the fewest number of animals?
- Which species that meet the above criteria are most economical to acquire and house?
Many model organisms have been shown to be particularly useful for certain types of genetic research. For example,
Mus musculus (House Mouse)
The mouse genome is similar to our own, so mouse genetic research is particularly useful for the study of human diseases. Mice are very useful for studying diseases such as atherosclerosis and hypertension. [3] They are better for studying complex biological systems found in humans such as the immune, endocrine, nervous, cardiovascular, and skeletal systems.
Caenorhabditis elegans (Nematode)
Studying cell death (apoptosis) in C. elegans could provide important information for counteracting effects of aging in humans as well as providing clues about cancer, diabetes and other diseases. [4]
Drosophila melanogaster (Fruitfly)
Drosophila are central to studies that explore human development, behavior, and genetic diseases. Genes associated with these attributes in humans have closely matched fly counterparts.
Saccharomyces cerevisiae (Yeast)
S. cerevisiae reproduces rapidly and in large quantities, making it an excellent model organism for studying metabolic processes.
Xenopus tropicalis (Western Clawed Frog)
Xenopus allows us to decipher how regulatory and interaction networks direct embryonic development and how they adapt during aging and under environmental stress. [5]
Mus musculus (House Mouse)
The mouse genome is similar to our own, so mouse genetic research is particularly useful for the study of human diseases. Mice are very useful for studying diseases such as atherosclerosis and hypertension. [3] They are better for studying complex biological systems found in humans such as the immune, endocrine, nervous, cardiovascular, and skeletal systems.
Caenorhabditis elegans (Nematode)
Studying cell death (apoptosis) in C. elegans could provide important information for counteracting effects of aging in humans as well as providing clues about cancer, diabetes and other diseases. [4]
Drosophila melanogaster (Fruitfly)
Drosophila are central to studies that explore human development, behavior, and genetic diseases. Genes associated with these attributes in humans have closely matched fly counterparts.
Saccharomyces cerevisiae (Yeast)
S. cerevisiae reproduces rapidly and in large quantities, making it an excellent model organism for studying metabolic processes.
Xenopus tropicalis (Western Clawed Frog)
Xenopus allows us to decipher how regulatory and interaction networks direct embryonic development and how they adapt during aging and under environmental stress. [5]
Using Model Organism Databases
Model Organism Databases are databases that consist of detailed biological data such as gene ontology, phenotypes, associated diseases, and expression. These databases are useful in evaluating similarities and differences between phenotypes across various model organisms. There are many different model organism databases such as:
Flybase
Flybase provided information regarding TCF4 phenotypes observed in drosophila. For example, expression of TCF4 under the control of GAL4 induces ectopic bristle formation on the thorax. [6] Expression of TCF4 under the control of two copies of GAL4 results in a rough eye phenotype. Constitutive expression of TCF4 under the control of GAL4 leads to embryo lethality.
Mouse Database (MGI)
MdI provided information regarding TCF4 phenotypes observed in Mus musculus. Homozygotes for a null allele show a partial block in early thymopoiesis (thymocyte differentiation). In addition, increased double-negative T cell count and increased sensitivity to anti-CD3 induced apoptosis is observed. [7] Homozygotes for another null allele show neonatal or postnatal lethality, reduced pro-B cell number, and abnormal pontine nuclei. Some types of phenotypes that attribute heavily to mutations/alleles of TCF4 are behavior/neurological, endocrine/exocrine glands, hematopoietic system, immune system, mortality/aging, nervous system, skeleton and vision/eye. [7]
Rat Database (RGD)
RGD provided information regarding TCF4 phenotypes observed in Rattus norvegicus. There were many different abnormalities, disabilities or diseases associated with the TCF4 gene in R. norvegicus. Some of these included symptoms very common for PTHS patients such as craniofacial abnormalities, developmental disabilities, neurodevelopment disorders, seizures and intellectual disability. However, the listed disease annotations also included diabetes, heart diseases, and liver cirrhosis. [8]
Flybase
Flybase provided information regarding TCF4 phenotypes observed in drosophila. For example, expression of TCF4 under the control of GAL4 induces ectopic bristle formation on the thorax. [6] Expression of TCF4 under the control of two copies of GAL4 results in a rough eye phenotype. Constitutive expression of TCF4 under the control of GAL4 leads to embryo lethality.
Mouse Database (MGI)
MdI provided information regarding TCF4 phenotypes observed in Mus musculus. Homozygotes for a null allele show a partial block in early thymopoiesis (thymocyte differentiation). In addition, increased double-negative T cell count and increased sensitivity to anti-CD3 induced apoptosis is observed. [7] Homozygotes for another null allele show neonatal or postnatal lethality, reduced pro-B cell number, and abnormal pontine nuclei. Some types of phenotypes that attribute heavily to mutations/alleles of TCF4 are behavior/neurological, endocrine/exocrine glands, hematopoietic system, immune system, mortality/aging, nervous system, skeleton and vision/eye. [7]
Rat Database (RGD)
RGD provided information regarding TCF4 phenotypes observed in Rattus norvegicus. There were many different abnormalities, disabilities or diseases associated with the TCF4 gene in R. norvegicus. Some of these included symptoms very common for PTHS patients such as craniofacial abnormalities, developmental disabilities, neurodevelopment disorders, seizures and intellectual disability. However, the listed disease annotations also included diabetes, heart diseases, and liver cirrhosis. [8]
What model organisms were used to study TCF4?
While browsing through various model organism databases, the Mouse Database (MGI) appeared to reveal phenotypes most similar to the phenotypes observed in humans. Hence, Mus musculus is the most appropriate model organism for this study of the TCF4 gene/protein.
Conclusion
Model organisms are crucial to the study of biological processes. There has been a vast amount of research conducted using model organism, such as those listed above, in order to more thoroughly understand the complex pathways and molecular players that contribute to process such as metabolism and neurological development. Although all model organisms are extremely helpful in research, certain organisms are more informative for certain types of research such as neurological processes or the study of metabolism. Due to the importance of TCF4 in neurological development and its important role as an E-protein/transcription factor, I narrowed my choice of model organism down to M. musculus, D. melanogaster, and C. elegans. Upon evaluating the appropriateness of each model organism for this study, I decided to use M. musculus, known as the house mouse. In addition to the high level of TCF4 conservation in M. musculus, previous research conducted on M. musculus supported my decision to use M. musculus as the model organism for this study. There have been various studies done regarding the role of TCF4 as a transcription factor/E-protein and its important interactions with other proteins for cellular processes. [9] This will allow me to evaluate the impacts of the TCF4 protein on cell differentiation processes.
References
Header: http://biochemistri.es/post/116597097236/e-coli-and-elephants
[1] YourGenome. (n.d.). What are model organisms? Retrieved from https://www.yourgenome.org/facts/what-are-model-organisms
[2] National Research Council (US) Committee on Dogs. Laboratory Animal Management: Dogs. Washington (DC): National Academies Press (US); 1994. 2, Criteria for Selecting Experimental Animals. Available from: https://www.ncbi.nlm.nih.gov/books/NBK236591/
[3] YourGenome. (n.d.). Why use the mouse in research? Retrieved March 1, 2018, from https://www.yourgenome.org/facts/why-use-the-mouse-in-research
[4] YourGenome. (n.d.). Why use the worm in research? Retrieved March 1, 2018, from https://www.yourgenome.org/facts/why-use-the-worm-in-research
[5] Xenopus. (n.d.). Introduction to Xenopus. Retrieved March 1, 2018, from http://www.xenbase.org/anatomy/intro.do
[6] Tamberg, L., Sepp, M., Timmusk, T., Palgi, M. (2015). Introducing Pitt-Hopkins syndrome-associated mutations of TCF4 to Drosophila daughterless. Biol. Open 4(12): 1762--1771.
[7] TCF4. (n.d.). MGI. Retrieved March 1, 2018, from http://www.informatics.jax.org/marker/MGI:98506
[8] RGD. (n.d.). Gene: Tcf4 (transcription factor 4) Rattus norvegicus. Retrieved March 1, 2018, from https://www.rgd.mcw.edu/rgdweb/report/gene/main.html?id=69271
[9] Flora, A., Garcia, J. J., Thaller, C., & Zoghbi, H. Y. (2007). The E-protein Tcf4 interacts with Math1 to regulate differentiation of a specific subset of neuronal progenitors. Proceedings of the National Academy of Sciences of the United States of America, 104(39), 15382–15387. http://doi.org/10.1073/pnas.0707456104
Figure 1. https://biology.uiowa.edu/model-organisms
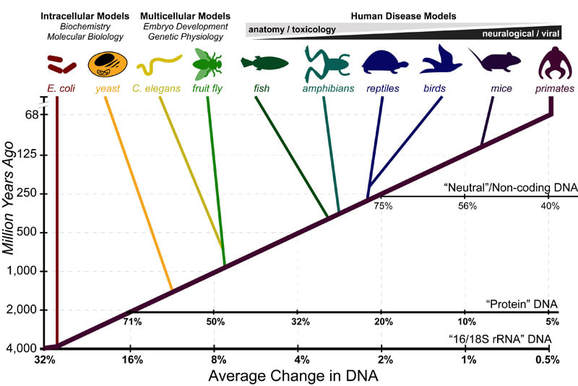
Figure 2. https://www.practicallyscience.com/model-organisms-and-dnas-molecular-clock/
Header: http://biochemistri.es/post/116597097236/e-coli-and-elephants
[1] YourGenome. (n.d.). What are model organisms? Retrieved from https://www.yourgenome.org/facts/what-are-model-organisms
[2] National Research Council (US) Committee on Dogs. Laboratory Animal Management: Dogs. Washington (DC): National Academies Press (US); 1994. 2, Criteria for Selecting Experimental Animals. Available from: https://www.ncbi.nlm.nih.gov/books/NBK236591/
[3] YourGenome. (n.d.). Why use the mouse in research? Retrieved March 1, 2018, from https://www.yourgenome.org/facts/why-use-the-mouse-in-research
[4] YourGenome. (n.d.). Why use the worm in research? Retrieved March 1, 2018, from https://www.yourgenome.org/facts/why-use-the-worm-in-research
[5] Xenopus. (n.d.). Introduction to Xenopus. Retrieved March 1, 2018, from http://www.xenbase.org/anatomy/intro.do
[6] Tamberg, L., Sepp, M., Timmusk, T., Palgi, M. (2015). Introducing Pitt-Hopkins syndrome-associated mutations of TCF4 to Drosophila daughterless. Biol. Open 4(12): 1762--1771.
[7] TCF4. (n.d.). MGI. Retrieved March 1, 2018, from http://www.informatics.jax.org/marker/MGI:98506
[8] RGD. (n.d.). Gene: Tcf4 (transcription factor 4) Rattus norvegicus. Retrieved March 1, 2018, from https://www.rgd.mcw.edu/rgdweb/report/gene/main.html?id=69271
[9] Flora, A., Garcia, J. J., Thaller, C., & Zoghbi, H. Y. (2007). The E-protein Tcf4 interacts with Math1 to regulate differentiation of a specific subset of neuronal progenitors. Proceedings of the National Academy of Sciences of the United States of America, 104(39), 15382–15387. http://doi.org/10.1073/pnas.0707456104
Figure 1. https://biology.uiowa.edu/model-organisms
Figure 2. https://www.practicallyscience.com/model-organisms-and-dnas-molecular-clock/
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.