What is Pitt-Hopkins Syndrome?
The first recorded cases of Pitt-Hopkins Syndrome (PTHS) occurred in 1978 at the Royal Children’s Hospital in Melbourne. A pediatrician, David Pitt, and pediatric neurologist, Ian Hopkins, described two unrelated patients with “severe intellectual disability who while awake had spells of rapid over-breathing followed by holding of breath until cyanosis.” [1] The patients had similar features such as wide mouth and palate, thick fleshy lips, broad beaked nose with flared nostrils and clubbing of fingers and toes. Throughout the following 28 years, only six new patients with the same syndrome were found in published work.
PTHS is a genetic disorder which affects a specific gene in chromosome 18, called TCF4. It is considered an Autism Spectrum disorder and some individuals diagnosed with PTHS have already been diagnosed with Autism, 'atypical' autistic characteristics or sensory integration dysfunction. [2] It is a very rare condition with approximately 500 affected individuals reported worldwide. [3]
PTHS is a genetic disorder which affects a specific gene in chromosome 18, called TCF4. It is considered an Autism Spectrum disorder and some individuals diagnosed with PTHS have already been diagnosed with Autism, 'atypical' autistic characteristics or sensory integration dysfunction. [2] It is a very rare condition with approximately 500 affected individuals reported worldwide. [3]
What causes Pitt-Hopkins Syndrome?
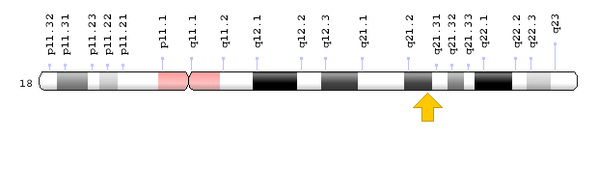
The cause of PTHS is de novo haploinsufficiency of the TCF4 gene, highlighted by the yellow arrow in Figure 1. [3] This means that individuals only have one functional copy of this gene and that this copy does not produce enough gene product to bring about the wild-type condition. PTHS is inherited in an autosomal dominant pattern, hence one copy of the gene mutation is sufficient to pass on the disorder. [3]
Gene and Protein Function
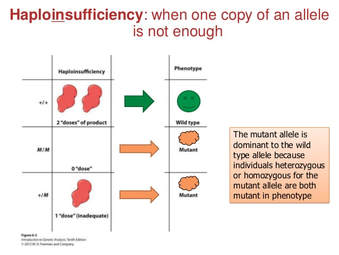 Fig 2. Mutation in TCF4 in patients with PTHS
Fig 2. Mutation in TCF4 in patients with PTHS
PTHS is caused by mutations in the TCF4 gene. Due to its DNA binding and gene controlling activities, the TCF4 protein is known as a transcription factor. Therefore, it is important to the maturation of cells to carry out various functions (cell differentiation) and the self-destruction of cells (apoptosis). [3] Mutations present in this gene disrupts the protein's ability to bind to DNA and control gene activity. [1] These disruptions, specifically controlling the activity of genes involved in nervous system development and function contribute to the characteristics and symptoms of PTHS. [3]
The TCF4 protein interacts with other proteins to carry out certain functions. However, when the TCF4 protein is nonfunctional, these other proteins are also affected. Lost of function in other proteins attached to the TCF4 proteins contribute to symptoms and conditions of PTHS. For example, the loss of the ASCL1 protein is believed to be responsible for breathing problems associated with PTHS. [3]
The TCF4 protein interacts with other proteins to carry out certain functions. However, when the TCF4 protein is nonfunctional, these other proteins are also affected. Lost of function in other proteins attached to the TCF4 proteins contribute to symptoms and conditions of PTHS. For example, the loss of the ASCL1 protein is believed to be responsible for breathing problems associated with PTHS. [3]
Symptoms of PTHS
|
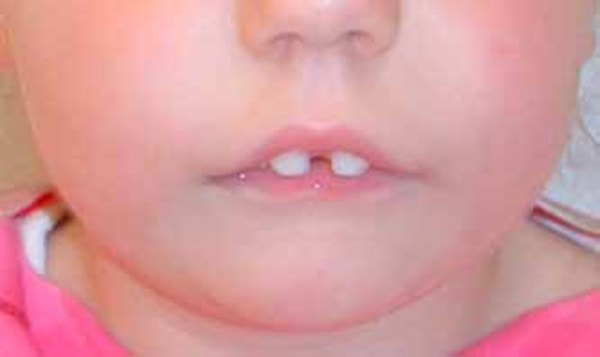
PTHS is characterized by: [3]
|
Epidemiology
Due to the infrequent diagnosis of PTHS, there are not significant population-based prevalence or incidence statistics. Until 2007, PTHS was not frequently reported, but molecular testing has increasing the number of patients reported. [1] There have been 80 patients confirmed with molecular genetic confirmation using array-Comparative Genomic Hybridization (CGH), mainly from Europe, the United States, and Canada. During a 17-month study period, Rosenfeld et al. screened 13,186 samples from intellectually disabled individuals. [5] Array-CGH was used and successfully found seven individuals carrying a deletion which included the whole or part of the TCF4 gene. [1] The authors estimated population frequency of PTHS caused by micro deletion in Washington to be 1/34,000-1/41,000. Actual prevalence is more frequently caused by point mutations, resulting in a higher true prevalence. [1]
Pitt Hopkins Research Foundation
200 Harmon Court
Winston-Salem, NC 27106 USA
Phone: 323-273-2632
Email: [email protected]
pitthopkins.org
200 Harmon Court
Winston-Salem, NC 27106 USA
Phone: 323-273-2632
Email: [email protected]
pitthopkins.org
References
Header: https://fineartamerica.com/featured/dna-dreaming-5-russell-kightley.html
[1] Peippo, M., & Ignatius, J. (2012, April). Pitt-Hopkins Syndrome. <https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3366706/>
[2] What is Pitt Hopkins syndrome? (2017, October 18). <https://pitthopkins.org/what-is-pitt-hopkins-syndrome/>
[3] Pitt-Hopkins syndrome - Genetics Home Reference. (n.d.). <https://ghr.nlm.nih.gov/condition/pitt-hopkins-syndrome>
[4] Micro Genomics. (2018). CGH-Array/Molecular Karyotype. < http://www.microgenomics.it/tecnologie/cariotipo-molecolare-array-cgh/?lang=en>
[5] Rosenfeld JA, Leppig K, Ballif BC, Thiese H, Erdie-Lalena C, et al. Genotype-phenotype analysis of TCF4 mutations causing Pitt-Hopkins syndrome shows increased seizure activity with missense mutations. Genet Med. 2009;11:797–805.
Figure 1. https://ghr.nlm.nih.gov/gene/TCF4
Figure 2. https://www.slideshare.net/vanessawhitehawk/genetics-chapter-4-part-1
Figure 3. https://ghr.nlm.nih.gov/art/large/exaggerated-cupids-bow.jpeg?ow
Header: https://fineartamerica.com/featured/dna-dreaming-5-russell-kightley.html
[1] Peippo, M., & Ignatius, J. (2012, April). Pitt-Hopkins Syndrome. <https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3366706/>
[2] What is Pitt Hopkins syndrome? (2017, October 18). <https://pitthopkins.org/what-is-pitt-hopkins-syndrome/>
[3] Pitt-Hopkins syndrome - Genetics Home Reference. (n.d.). <https://ghr.nlm.nih.gov/condition/pitt-hopkins-syndrome>
[4] Micro Genomics. (2018). CGH-Array/Molecular Karyotype. < http://www.microgenomics.it/tecnologie/cariotipo-molecolare-array-cgh/?lang=en>
[5] Rosenfeld JA, Leppig K, Ballif BC, Thiese H, Erdie-Lalena C, et al. Genotype-phenotype analysis of TCF4 mutations causing Pitt-Hopkins syndrome shows increased seizure activity with missense mutations. Genet Med. 2009;11:797–805.
Figure 1. https://ghr.nlm.nih.gov/gene/TCF4
Figure 2. https://www.slideshare.net/vanessawhitehawk/genetics-chapter-4-part-1
Figure 3. https://ghr.nlm.nih.gov/art/large/exaggerated-cupids-bow.jpeg?ow